Bioactivity descriptors for uncharacterized compounds
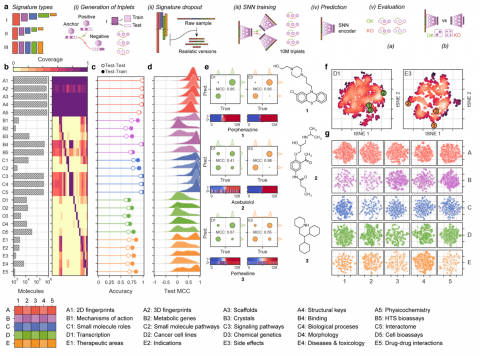
Bertoni M, Duran-Frigola M, Badia-i-Mompel P, Orozco-Ruiz M, Guitart-Pla O, Aloy P, Chemical descriptors encode the physicochemical and structural properties of small molecules, and they are at the core of chemoinformatics. The broad release of bioactivity data has prompted enriched representations of compounds, reaching beyond chemical structures and capturing their known biological properties. Unfortunately, bioactivity descriptors are not available for most small molecules, which limits their applicability to a few thousand well characterized compounds. Here we present a collection of deep neural networks able to infer bioactivity signatures for any compound of interest, even when little or no experimental information is available for them. Our signaturizers relate to bioactivities of 25 different types (including target profiles, cellular response and clinical outcomes) and can be used as drop-in replacements for chemical descriptors in day-to-day chemoinformatics tasks. Indeed, we illustrate how inferred bioactivity signatures are useful to navigate the chemical space in a biologically relevant manner, and unveil higher-order organization in drugs and natural product collections. Moreover, we implement a battery of signature-activity relationship (SigAR) models and show a substantial improvement in performance, with respect to chemistry-based classifiers, across a series of biophysics and physiology activity prediction benchmarks.
Nature Communications,
2021, 12, 1-13
Pubmed: 34168145
Direct link: 10.1038/s41467-021-24150-4
